Experts in typing bacterial strains by whole genome sequencing
Fully characterise your bacterial strain with our typing service
The increasing global demand and use of microbial biocontrol agents and biofertilisers have prompted efforts in screening for candidate bacterial strains with activity against soil-borne pathogens and/or plant-growth promotion capabilities. Requirements for the registration of microbial-based products include, among others, a precise identification of the bacterial strain, as well as data on the efficacy and safety of the inoculum. Whole-genome sequencing (WGS) has become the gold standard approach for identifying bacteria at the strain level. WGS is a fast and cost-efficient way to generate high-resolution genomic data, which brings a number of advantageous applications for the biocontrol industry, including:
- Complete characterisation of bacterial strains, to meet the requirements for product registration and patent application.
- Accurate discrimination of closely related and/or commercial strains.
- Identification of traits of agronomical interest (e.g., genes related to biological nitrogen fixation, phosphate solubilisation, ACC deaminase activity, and production of siderophores and phytohormones; chemotaxis and flagellar motility genes; secondary metabolic pathways).
Our bacterial strain typing service uses high-throughput sequencing to obtain the annotated genomes of your bacterial strains. To do so we take advantage of Illumina’s MiSeq and NovaSeq technologies which enable us to obtain the genomes cost-effectively and accurately.
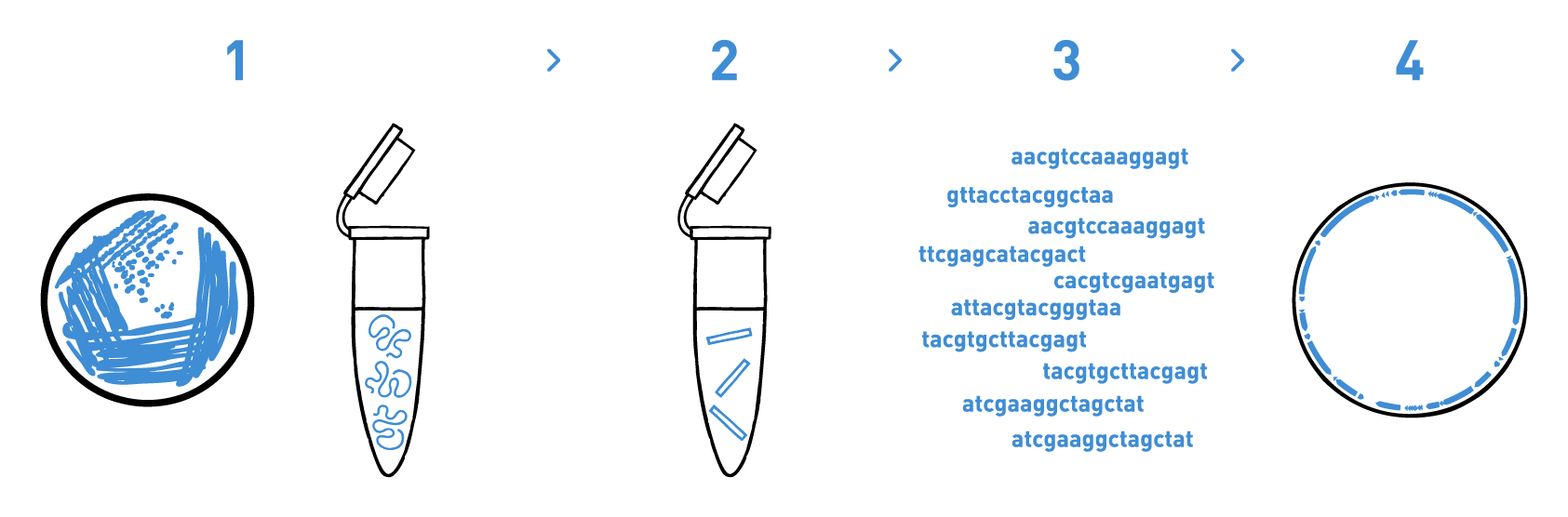
Step 1
We isolate DNA from your samples. We have adapted different DNA isolation protocols, depending on the starting biological material.
Step 2
We prepare genomic libraries using the kits recommended by Illumina.
Step 3
We sequence the libraries using Illumina's MiSeq or NovaSeq technologies. Millions of reads per library are obtained in this step.
Step 4
We analyse the high-throughput sequencing data to obtain the annotated genomes of your bacterial strains.
What your receive
What we need
Your bacterial cultures appropriately preserved (fresh, frozen, or lyophilised). Please contact us if you need assistance on how to preserve and ship your cultures.