Experts in microsatellite development
Microsatellite markers for your species with our microsatellite development service
AllGenetics' microsatellite development service uses high-throughput sequencing to obtain primer pairs which amplify polymorphic microsatellite loci in your study species. We multiplex and test for polymorphism the primers obtained in a number of individuals from different populations.
Our team has expertise in developing microsatellite markers for either animal, plant, and fungal species. We are also trained in genotyping individuals from polyploid populations.
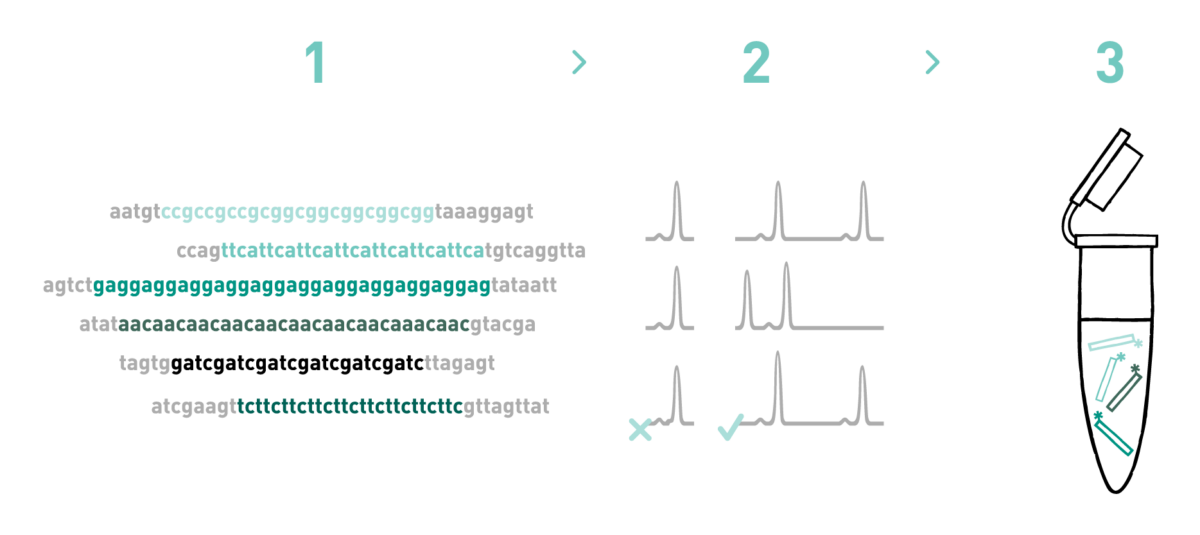
Step 1
We generate a microsatellite-enriched genomic library. To do so we use high-quality genomic DNA from 1 individual of your study species. Then, we sequence this library using the Illumina NovaSeq platform and obtain thousands of microsatellite-containing reads. Finally, based on the microsatellite-containing reads obtained previously, we design 500 primer pairs on the flanking regions of microsatellite motifs.
Step 2
Primer pairs will be multiplexed in sets of 3 - 5, based on their specific features. These primers will be tested in a number of individuals. You decide how many primer pairs you would like us to test and the number of individuals you would like us to genotype. Even though the number of primer pairs amplifying polymorphic loci is extremely dependent on the biological features of the species under study, as a general rule we suggest to test 4 times as many primer pairs as polymorphic loci you need for your project.
Step 3
We select only those primer pairs which amplified polymorphic loci in Step 2 and rearrange the multiplexes in order to optimise them in silico.
For your convenience, we can carry out the entire project, step 1 alone, or steps 1+2.
What you receive

Steps 1+2: The deliverables of step 1 + the PCR protocol and the multiplex organisation, the results of the biological test, and the primers and fluorescent probes (shipment usually restricted to the European Union).
Steps 1+2+3: Everything in step 1 and step 2 + the in silico optimised multiplex setup.