Experts in bacterial genome sequencing
Characterise your bacterial strains through our whole genome sequencing service
We are eager to tailor our protocols to the needs of your research project: therefore, we can take advantage of both short-read (Illumina) or long-read (PacBio and Oxford Nanopore) sequencing technologies, and perform on-demand, custom bioinformatic analyses.
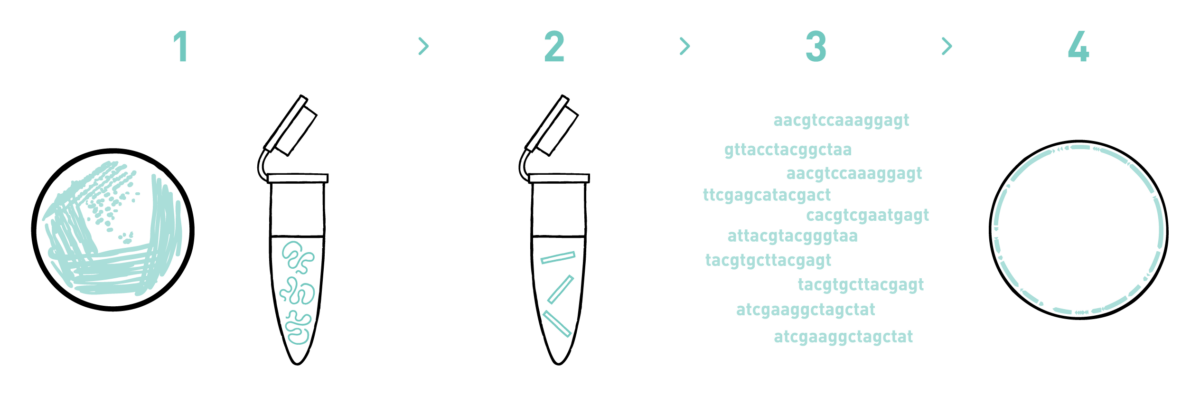
Step 1
We isolate DNA from your samples using specific DNA isolation protocols for bacterial cells.
Step 2
We prepare genomic libraries using the kits recommended by Illumina.
Step 3
We sequence the libraries using Illumina's MiSeq or NovaSeq technologies. As a norm, we obtain 500 megabases of raw data per library but we can adapt the output to the genome size of the species under study.
Step 4
We analyse the high-throughput sequencing data to obtain your annotated genomes.
What you receive
A report with a summary of the methods followed.

The raw high-throughput sequencing files, which will be delivered through our server, and the corresponding quality control report including demultiplexing and pre-processing.