Experts in MeRIP-seq
Identification of m6A modifications in the RNA with AllGenetics' MeRIP-seq service
With our MeRIP-seq service (also known as m6A-seq) we identify N6-methyladenosine (m6A) modifications in the RNA. These modifications are known to have a large effect on gene expression and mRNA metabolism.
Our lab team immunoprecipitates the RNA containing methylated adenosines by using a specific antibody. We then use the immunoprecipitated RNA for library construction, and we sequence it together with a control RNA sample.
Our bioinformatics team map the reads coming from both the immunoprecipitate sample and the control to a reference transcriptome and compare them to find regions of differential coverage. These regions will point to the location of the m6A residues.
Our full MeRIP-seq service includes RNA fragmentation, RNA immunoprecipitation, library construction of both the immunoprecipitate and the control RNA, high-throughput sequencing, and bioinformatic analyses.
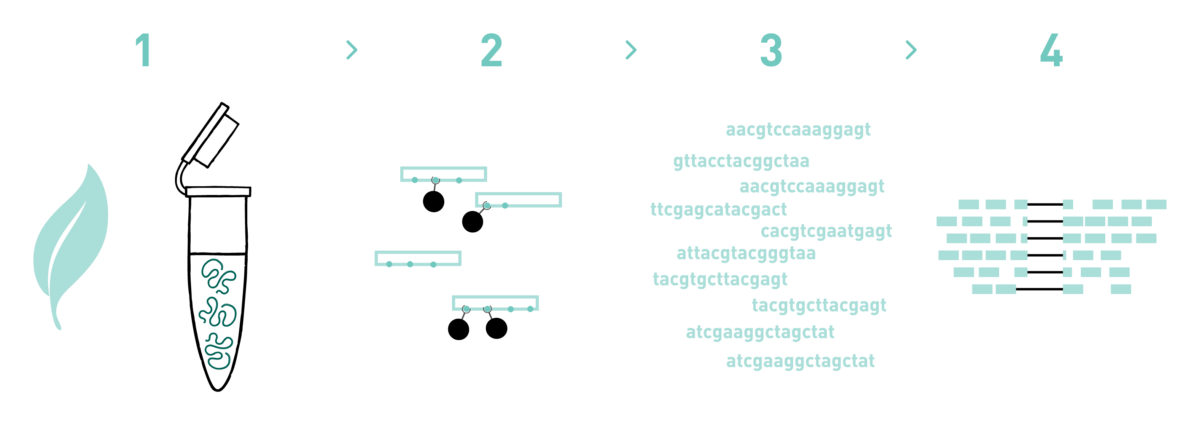
Step 1
We isolate RNA from your samples. We have adapted different RNA isolation protocols, depending on the starting biological material.
Step 2
We inmunoprecipitate your samples and prepare cDNA libraries compatible with Illumina’s sequencing platforms.
Step 3
We sequence the libraries using the Illumina NovaSeq technology. Millions of reads per library are obtained in this step.
Step 4
We analyse the high-throughput sequencing data obtained in the previous step.
For your convenience, we can carry out the entire project or only those steps you require. As a norm, steps 2 and 3 should be ordered jointly.
What you receive

The raw high-throughput sequencing files, which will be delivered through our server, and the corresponding quality control report including demultiplexing and pre-processing.