Experts in typing fungal strains by whole genome sequencing
Fungal strain typing service using whole genome sequencing
Arbuscular mycorrhizal (AM) fungi and antagonistic fungi, such as Trichoderma spp. or Chaetomium spp., are well-known beneficial microorganisms used as biocontrol agents and/or biofertilisers. Efficient biocontrol or plant growth-promoting fungal strains are usually selected among numerous, sometimes closely related, less effective strains.
The development of commercial fungal-based biocontrol products and biofertilisers can greatly benefit from recent advances in genomic analysis. The staggering amount of information provided by whole genome sequencing (WGS) and comparative genomic studies of fungal strains now opens up the possibility to better understand the mechanisms underlying their antagonistic and plant growth promotion activities. This, in turn, will aid in the identification and selection of fungal strains with improved biocontrol and/or biofertiliser potential. Some common applications of our fungal strain typing service include:
- A complete characterisation of fungal strains, to meet the requirements for product registration and patent application.
- Accurate discrimination of closely related and/or commercial strains.
- Development of a molecular identification method to monitor the persistence and distribution of a particular strain in the field.
- Screening for relevant genes involved in traits of agronomic interest (e.g., virulence factors and pathogenicity genes; genes involved in nutrient uptake and solubilisation; candidate secreted effector proteins; or secondary metabolism gene clusters involved in the synthesis of antifungal compounds).
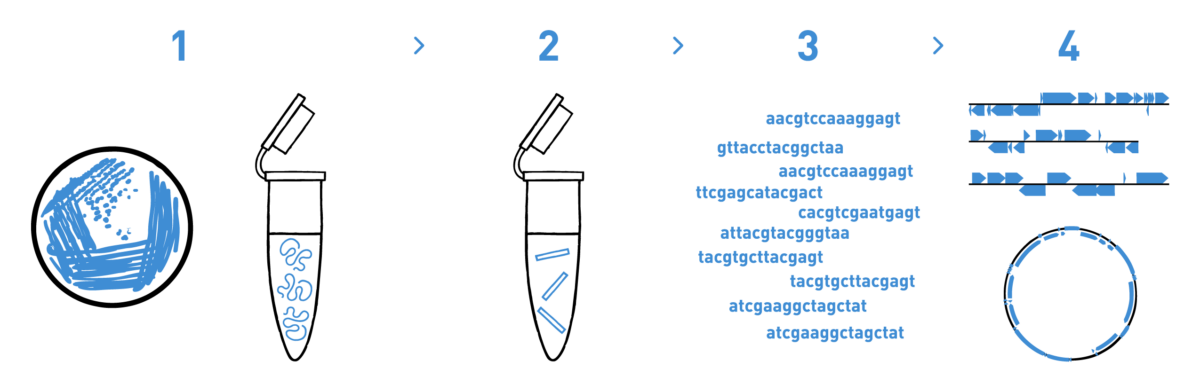
Step 1
We isolate DNA from your samples. We have adapted different DNA isolation protocols, depending on the starting biological material.
Step 2
We prepare genomic libraries using the kits recommended by Illumina.
Step 3
We sequence the libraries using the Illumina NovaSeq technology.
Step 4
We analyse the high-throughput sequencing data to obtain your annotated contigs.
What you receive
- Automatic gene annotation.
- Average nucleotide identity between the genomes sequenced and other yeast genomes available in reference databases.
What we need
Your fungal cultures appropriately preserved (fresh, frozen, or lyophilised). Please contact us if you need assistance on how to preserve and ship your cultures.