Experts in characterising vineyard and wine microbiome by DNA metabarcoding
Vineyard and wine microbial diversity characterisation service
Microbial communities are a vital part of agrarian soils, which play an important role in grapevine growth and development, contributing to the terroir. In the last few years, more and more studies link the microbial community structure in vineyards and cellars to variations in the organoleptic properties of wine.
High-throughput sequencing (HTS) technologies have revolutionised how microbial communities are studied, which allows for a precise characterisation of all the species present in a given sample, as long as reference sequences are available. Even those species which cannot be detected by culture-dependent methods can be identified.
Our vineyard and wine microbial diversity characterisation service uses DNA metabarcoding to screen the bacterial, fungal, protist, and archaea species present in your samples, whatever their source might be (soil, plant tissues, must, or wine).
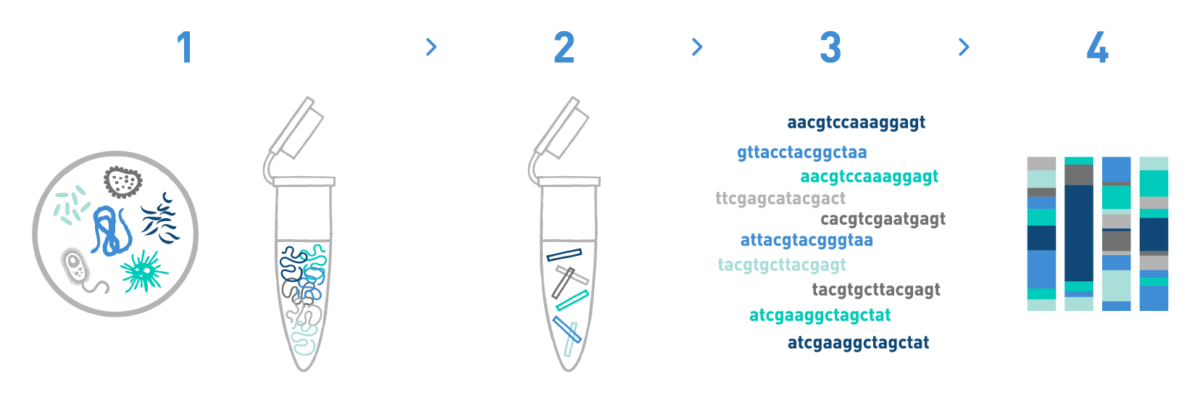
Step 1
We isolate DNA from your soil or plant samples. We have adapted different DNA isolation protocols, depending on the starting biological material.
Step 2
We prepare DNA metabarcoding libraries for the target taxonomic group(s) -bacteria, archaea, protists, and/or fungi-. For this we use universal or in-house-developed primer pairs.
Step 3
We sequence the DNA metabarcoding libraries using Illumina's MiSeq or NovaSeq technologies.
Section 4
We analyse the data obtained in the previous step to get the taxonomic composition of the samples.
What you receive
- A list of the Operational Taxonomic Units (OTUs) retrieved per sample.
- The number of sequence reads assigned to each OTU in each sample.
- A graphic representation of the relative abundances of the different OTUs in each sample.
What we need
Your samples which have been appropriately preserved (ethanol, silica gel, frozen, or another suitable preservation method). If required, we provide sampling kits and sampling collection guidelines to ensure that your samples arrive at our lab in optimal conditions.