Experts in identifying grapevine varieties by microsatellite markers
Grapevine varieties identification service using microsatellite markers
For over the past two decades, microsatellites or simple sequence repeats (SSRs) have been the most widely used molecular markers in plant genetic studies. In the particular case of grapevines, microsatellites have been extensively used for cultivar or varietal DNA fingerprinting, as recommended by the International Union for the Protection of New Varieties of Plants (UPOV 2010, 2011).
Our grapevine varieties identification service uses the following internationally accepted loci: VVS2, VVMD5, VVMD7, VVMD25, VVMD27, VVMD28, VVMD32, VrZAG62, and VrZAG79. This analysis will allow you to identify and verify your varieties and cultivars for breeding programs or legal protection. They will also serve as a cross-reference for authorised varieties.
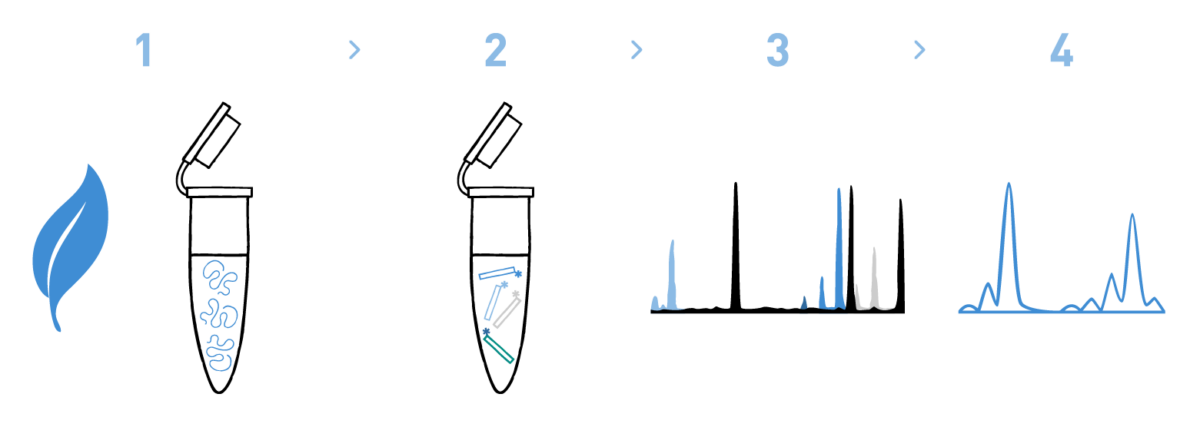
Step 1
We isolate DNA from your grapevine samples.
Step 2
We amplify the microsatellite loci under study using fluorescently-labelled primer pairs.
Step 3
We run the labelled amplicons in the sequencer.
Step 4
We analyse the electropherograms and call the alleles observed at each locus.
What you receive
What we need
Your tissue samples which have been appropriately preserved (ethanol, silica gel, frozen, or another suitable preservation method). If required, we provide sampling kits and sampling collection guidelines to ensure that your samples arrive at our lab in optimal conditions.