Experts in phage genome sequencing
Phage characterisation service through whole genome sequencing
With AllGenetics' phage genome sequencing service you will obtain the annotated genomes of your phages. We will purify phage DNA or RNA (either single-stranded or double-stranded) using specific nucleic acid isolation protocols. We will delete the remaining bacterial contamination bioinformatically after sequencing.
With this service you will be able to analyse isolated phages. If your samples contain a viral mix, please consider our metagenomics and metatranscriptomics services.
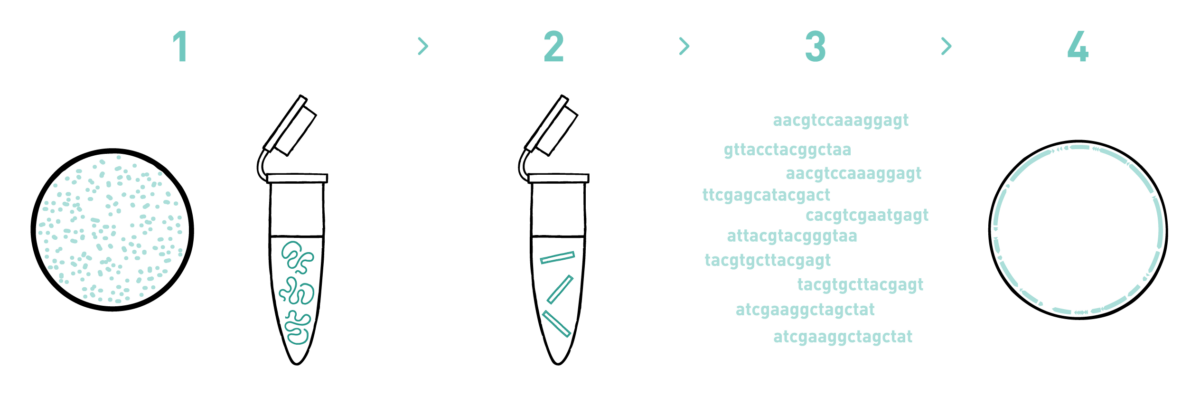
Step 1
We isolate DNA or RNA from your purified phage samples using a phage nucleic acid isolation protocol.
Step 2
We prepare genomic libraries using a combination of specific in-house developed protocols and commercial kits suitable for Illumina sequencing.
Step 3
We sequence the libraries using the Illumina MiSeq technology. As a norm, we obtain 5 megabases of raw data per library but we can adapt the output to the genome size of the phage under study.
Step 4
We analyse the high-throughput sequencing data to obtain your annotated genomes.